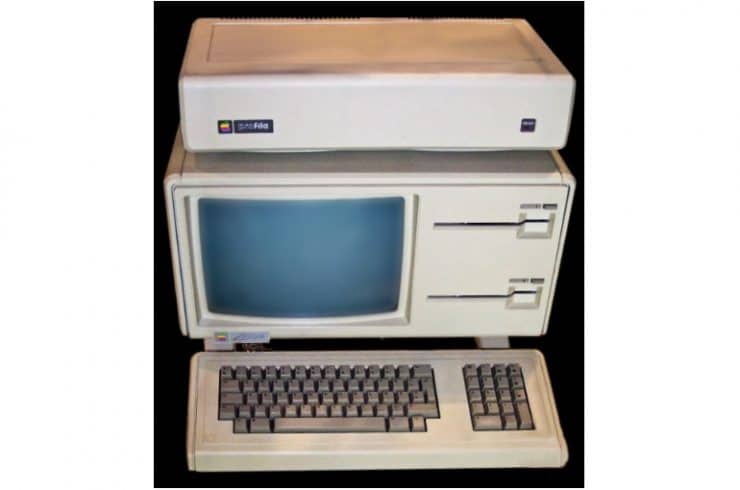
Advances in DNA technology bring up fascinating questions about what role it will play in our society, from medicine to food
An arrest in the decades-old Golden State Killer case.
A Chinese scientist creating the first gene-edited twin baby girls.
DNA is clearly changing our reality.
In recognition of National DNA Day on April 25, scientists at Arizona State University took time to reflect on some big questions: What brought us to this point, where are we going from here — and just because we can, should we?
As is the case with most dense subjects, the best place to start is usually the beginning.
Where it all began
The average science novice might point to the Human Genome Project that had roots in the 1980s as the origin of modern DNA science. But it goes back further than that, to the discovery of the double helical structure in the 1950s and the development of the sequencing process in the 1970s that unlocked the genetic information contained in DNA.
“Those were crucial technological breakthroughs that enabled the whole field to unfold,” said Robert Cook-Deegan, professor in the School for the Future of Innovation in Society.
He witnessed firsthand as genomics took on its current form in the late 1980s, when molecular biologist James Watson — the very man who in 1953 had co-authored the paper proposing the double helix structure of the DNA molecule — asked him to lend his science and health policy expertise to the Human Genome Project.
At the time, computing technology began advancing at a rapid clip, allowing scientists to study the whole genome at once instead of one gene at a time — for the first time, they had a 30,000-foot view of the building blocks of life.
The term genomics was coined with the launch of the eponymous peer-reviewed journal in 1987 and helped to distinguish the science from genetics, the study of inheritance that only considered one gene at a time.
This newfound perspective of the curious interactions and fascinating entanglements of the chromosomes and proteins that make us who we are ushered in an era of more precise diagnostics. By analyzing a person’s genome and comparing it to relatives, scientists could pinpoint differences and similarities in their genetic makeup that might make them more prone to certain diseases or conditions.
“We’re all mountains, but we have some differences.”
— School of Life Sciences Assistant Professor Melissa Wilson
School of Life Sciences Assistant Professor Melissa Wilson studies the evolution of sex chromosomes and how they could be related to disease risk. In an unprecedented upcoming paper, she and a team of researchers theorize that women’s propensity toward overactive immune systems helps them both surveil and fight off cancer better than men.
She explains the utility of the human genome reference thusly:
“It’s like if I gave you a puzzle of Camelback Mountain and I said, ‘This is the human genome, it’s Camelback Mountain.’ But really, some of us look like the Appalachians, and some of us look like the Superstitions, and some of us look like Four Peaks. We’re all mountains, but we have some differences. So we use that puzzle of Camelback Mountain as our reference to see where they are the same and where they are different.”
Then, in the mid-2000s, new forms of faster DNA sequencing allowed for the detection of variants in individuals and populations.

Robert Cook-Deegan
“That’s one thing nobody saw coming,” Cook-Deegan said. The ability to identify genetic differences among populations has vast implications for tracing ancestry, including the study of ancient DNA. It gave researchers insight into regional ancestry, migration patterns and more.
Nowadays, while scientists have already harnessed the potential of the naturally occurring genome editing system known as CRISPR-Cas9 to genetically modify babies in the womb, Cook-Deegan cautions we still have much more to learn.
“We’re at the toddler stage,” he said. “There’s just so much data coming out and we know so little about so much. Understanding the genome is not just about what genes you have, but understanding why and how and when they’re turned on and turned off. … We still don’t understand that regulatory switch-work at all. We’re just at the very beginning of being able to understand that. That’s going to go on for about another century.”
The genome guides precision medicine
From the 18th through the 20th centuries, a physician’s dominant tool was the microscope. They would look at cells or tissues under a microscope and then say, “This patient has disease X, Y or Z,” based on the way the cells appeared. It was very good, and took health care a long way.
Then the Human Genome Project launched. The world’s largest collaborative biological project, it was an international scientific research project with the goal of determining the sequence of human DNA and identifying and mapping all of the genes of the human genome from a physical and a functional standpoint. It was completed in 2003.
“What we learned in the 21st century, or even at the very tail end of the 20th century, is that we can get even more precise about what a patient has by looking at the molecules,” said Joshua LaBaer, executive director of ASU’s Biodesign Institute and a professor in the School of Molecular Sciences. LaBaer is one of the nation’s foremost investigators in the rapidly expanding field of personalized diagnostics.
“Precision medicine is basically a way of fine-tuning the way we treat our patients,” LaBaer said. “With personalized medicine, doctors like myself always felt we personalized treatment. We don’t treat a population; we treat an individual.”
When LaBaer went to medical school back in the 20th century, one would look at certain cells and tissues in the breast under the microscope and say “infiltrating ductal carcinoma of the breast.” That was a pathologist’s terminology for breast cancer. Now doctors know that one disease under a microscope is like seven or eight different molecular diseases if you look more deeply. There’s luminal A type, luminal B type, HER2 type, there’s triple negative type, and so on. And those different types behave differently with different chemotherapies. They also respond to specific therapies that are not available for the others. And that’s just breast cancer. The same kinds of things are true for other types of cancers as well as other diseases.
“In the 21st century, we’re looking more at these molecules and we’re understanding much more about how they contribute to disease, what they tell us about the prognosis of the patient, and what opportunities of therapy we can bring to bear,” LaBaer said.
The Human Genome Project, for the first time, outlined a complete human parts list. Looking at the human genome basically told us all the different genes that are there. That was the first step, and it was a big one. But that project looked at a few people’s genomes, and people vary widely.
The All of Us Research Program was launched by the U.S. government in 2018. It seeks to extend precision medicine to all diseases by building a national research cohort of 1 million or more U.S. participants. Anyone over the age of 18 living in the United States can join.
We all have a likelihood of getting different diseases. But when we do, our outcomes can differ from person to person with the same disease. Much of it is a product of our different genomes.
“How do we understand the variation?” LaBaer said. “What is the variation between us, and how does understanding that variation help predict risks of disease and/or responses to disease when they occur? By cataloging all that information, we will learn a lot about those sorts of factors. That’s what (All of Us) does for us.”
There are limits to what genome info can do for disease risk. LaBaer’s favorite metaphor is the genome is a recipe, but people given the same recipe might make dishes that taste a little bit different. 
“The genome is the starting point, but it’s not the answer to everything.”
— Joshua LaBaer, professor and executive director of ASU’s Biodesign Institute.
The genome is the blueprint for how to make a person. People are a little different from the genome, because wear and tear happen to them. Things break. Sometimes people break even when they’ve always appeared to be fine, like a vegan athlete who develops diabetes in his late 40s.
“The genome doesn’t necessarily tell us what’s going to happen to a person,” LaBaer said. “It gives us the mathematical possibility of things that might happen to that person. … The genome can tell us likelihoods of our being able to metabolize certain drugs in certain ways. … That’s called pharmacogenomics, and that’s very important. The genome is the starting point, but it’s not the answer to everything.”
There are a lot of things about DNA information people need to know, LaBaer said. Although your entire human genome can be sequenced, fairly little is known about how to interpret that.
“If anyone tells you, ‘Oh, we’ll sequence your genome and that will fix everything,’ that’s probably not true,” he said. “It’s almost certainly not true. Certainly some of those elements are helpful. There are known genetic disorders you can detect.”
Whether you’re going to get heart disease or a specific type of cancer, mostly what’s now known can’t predict that. And, contrary to what you see on TV, genome sequencing can’t tell you whether your heritage is Albanian or Latvian. What do consumers need to watch out for?
“You need to be careful about what kind of promises are made about what you’re going to learn from this,” LaBaer said. “A lot of these companies initially promised all this medical value for people, and the FDA forced them to back away from that claim. Now most of them are marketing themselves as talking about your heritage. Even there, I think a lot of what’s promised is a little bit oversold at this point. When people say you’re 30 percent this and 15 percent that, I don’t know what that means. I don’t know how well that’s understood at this point. … DNA is only useful if the clinical information attached to it is also accurate. Oftentimes it isn’t.”
LaBaer cautions it’s worth looking at the fine print for privacy issues. Some of the companies sequencing genomes are selling that information to other companies for research purposes. Theoretically it’s not identified as yours. They’ll say it’s from a Caucasian female in her 30s, or something along those lines. A lot of their business models aren’t based on the fees you paid, but fees from selling the sequence to someone else. And, as is discussed in other sections of this series, there are no legal barriers from law enforcement going in to any of these companies and seeing what they have.
Finding solutions with gene therapies
When the gene editing tool CRISPR burst upon the scene in 2012, scientists immediately saw its potential to cure genetic diseases. Samira Kiani has built her career around her passion for applying CRISPR technology to synthetic biology. An assistant professor in the School of Biological and Health Systems Engineering, she has established her research program to combine CRISPR technology with synthetic biology to develop safer and controllable gene therapies.

Samira Kiani
Is that potential realistic? How viable are solutions?
There are three major areas CRISPR can potentially make an impact, according to Kiani. The first is gene therapy: Patients with formal genetic diseases like metabolic diseases or immune disorders have some sort of faulty genes.
“We can use CRISPR to disrupt those faulty genes or correct those faulty genes,” Kiani said. “This time CRISPR would allow us to pinpoint the type of genes that already exist in human DNA and just modify those, correct those or disrupt the faulty genes.”
Another potential arena for CRISPR would lie in correcting susceptibility genes that put people at risk of diseases like diabetes, cancer and atherosclerosis. A delivery device would put CRISPR in the patient’s body. The tool would go to a certain organ and change the genes.
“CRISPR would allow us at some point — let’s say five or 10 years from now — to develop a form of gene therapy using CRISPR and go and modulate those genes so that they are not really conferring susceptibility anymore to those diseases,” Kiani said.
The third application for human health Kiani cites is correcting a faulty gene at the embryonic level. For example, if a couple had genes that would immediately lead to a fetal disease, they could do in vitro fertilization and the genes could be corrected at the level of the embryo. Then the corrected embryo could be implanted.
CRISPR also is being used to diagnose certain genetic diseases or viruses that can infect cells such as HPV, HIV or Ebola.
Clinical applications are feasible within five to 10 years, according to Kiani. The technology is moving rapidly — but there’s a catch.
Science fiction writer William Gibson famously said, “The future is here. It’s just not widely distributed yet.” Travel from a big city to a rural town, or from an industrialized nation to a developing one, and unequal distribution of advanced anything is obvious.
“With technologies like this, you will face all the issues with access and equality of access,” Kiani said. “How do we make it affordable for every doctor’s office to have it? If we are speaking with regard to accessibility to patients at every doctor’s office, I would say a longer term — maybe 15 or 20 years. As any new technology is developed — internet technology or iPhone — every time these new technologies develop, rich (people) have better access to it. So I would say once this technology is rapidly developed, it’s either accessible to people with more money or governments and insurance companies need to come on board so they actually provide this accessibility to patients.”
Spinal muscular atrophy is a debilitating, muscle-wasting disease caused by death of nerve cells in the spine. The FDA approved the sale of a new drug for the treatment of this disease. The drug tricks the spinal neurons into using another gene to produce protein, allowing the patient to survive. Here’s the catch: The drug costs $750,000 in the first year followed by $375,000 a year after that — for life.
Gene therapies have the potential to alleviate that problem of cost. They require the creation of a drug specific for each patient. It has to be designed, customized, administered and monitored by several expert personnel. Currently, none of that comes cheap.
But there is a light at the end of that tunnel, Kiani said.
“The claim with CRISPR is because it’s easier to repurpose, the costs might be lower,” she said.
We can — but should we?
Ethical questions concerning biotechnology were already a part of the science and health policy conversation by the time the field of human genetics took off, thanks in part to biological weapons research that lasted until the Biological Weapons Convention in 1972 and the advent of agricultural biotechnology (which remains controversial to this day).
In relation to DNA science, School of Life Sciences Associate Professor Ben Hurlbut said ethical concerns arose out of the combination of the hopes that were attached to what knowledge the human genome could give us — such as the capacity to treat disease — and the uses it might be put to that could be contrary to the public good.
Hurlbut and colleagues are working on creating a new kind of structure for governance of the field — a global observatory for gene editing, which he wrote about in a March 2018 article for Nature.
“In the earliest days of the development of genetics and the technology associated with it, there was a tendency in the scientific community to ask those large ethical questions,” he said. “But over the years, there’s been a kind of resistance to that and a silencing of discussions that look far ahead.”
Cook-Deegan can attest to the former. A few years into working on the Human Genome Project, he authored “The Gene Wars: Science, Politics, and the Human Genome,” a personal account of the genesis and early stages of the project that also addressed anxieties regarding far-reaching medical and social implications. Later, he would go on to found Duke University’s Center for Genome Ethics, Law and Policy.
What is interesting about the field of human genetics, he noted, is that it started to take off at the same time that historians around the world were beginning to re-examine the history of eugenics and so-called “racial hygiene” that led to sterilization and interracial marriage bans. So as the field advanced, so too did unease about such ills resurfacing.
At the same time, most understood the potential health benefits of genomics.
“So from the beginning, there were ethical discussions and a parallel effort to do something about policy, to think about the legal issues that were going to need to be addressed,” Cook-Deegan said.
Some of the earliest ethical concerns with biotechnology were related to biosafety, military and industrial control of life and genetic engineering. Lately, as Hurlbut mentioned, things have become even more complicated. 
“Our ability to do stuff far exceeds our ability to do it ethically.”
— Andrew Maynard, professor in the School for the Future of Innovation in Society
In 2013, in response to a molecular diagnostic company that attempted to do so, the Supreme Court ruled that isolated human genes could not be patented. While proponents of the argument claimed patents would encourage investment in biotechnology and promote innovation in genetic research, opponents claimed patenting isolated genes would hamper further disease research and limit options for patients seeking genetic testing.
And there’s also reason to question whether we rely too much on what DNA tells us about disease risk factors to determine treatments and predict health outcomes.
“I’m not an MD,” Wilson said, “but for example, aspirin is advised to give to everyone to help prevent stroke. Turns out, it doesn’t really work in women. And this has been known for decades. But we just give it to them anyway.
“So we have personalized medicine based on populations that are not representative of the people we’re working on. If we really want to have personalized medicine, we need to actually have our data sets be representative of everyone. And they’re not right now, unfortunately.”
Andrew Maynard, professor in the School for the Future of Innovation in Society, studies emerging tech and responsible innovation. In his new book, “Films from the Future,” he grapples with a number of issues around the ethics of how we work with DNA and what it means to innovate responsibly.
In the years to come, he believes there is a growing urgency for not just scientists but everyone DNA technology has the potential to affect to learn how to be socially responsible with it.
“Our ability to do stuff far exceeds our ability to do it ethically,” he said. “So there’s a huge obligation for us to think critically about what we’re doing and have an open conversation about it.”
Gene modification on our tables
As for that controversial agricultural biotechnology, genetically modified organisms have been around since the early 1970s. Definitions vary, but consensus hovers around an organism that has been altered in a way that would not occur in nature.
A bacteria was the first organism to have its DNA altered, followed by a mouse and a plant. The first organism engineered for commercial ends was the Flavr Savr tomato, which hit supermarket shelves in 1994. The FDA declared it as safe as a natural tomato. The goal of all tomato growers is to be able to handle them as soon as possible and for them to have a longer shelf life. The maker’s intent was to slow down ripening. Flavr Savrs did have a longer shelf life, but they still had to be picked and handled like any vine-ripened tomato. The company struggled with profits, mainly because they didn’t know enough about the farming end of the business, and were eventually acquired by Monsanto.
Flash-forward another decade and GloFish hit the market. They’re still around, for people who think tropical fish are too drab. In 2015, AquAdvantage Atlantic salmon hit Canadian markets. Modified to grow to market size in 16 to 18 months instead of three years, it was initially blocked from being sold in the U.S. In early March, however, the FDA lifted the import ban on genetically engineered salmon and salmon eggs.
Oya Yazgan is a molecular biologist in the College of Integrative Sciences and Arts, where she teaches a course in food and human health. How foods are produced and the consequences of consuming various types of foods is her passion.
There’s one big question hovering over GMO foods: Are they safe? The short answer — no one really knows. Research has been done and used as a reference for saying that GMOs are safe, but it’s neither serious nor reliable science, Yazgan said.
“We need to take a very careful look at these before we play with people’s health.”
— Oya Yazgan, molecular biologist in the College of Integrative Sciences and Arts
“The studies they refer to are poorly designed and statistical analyses are not strong, and they are making conclusions that are not scientifically valid,” she said. “We have some preliminary evidence that needs stronger scientific research that indicates there are damages that are being caused by these GMOs. They are seeing intestinal damage in mice and pigs. The general bigger problem I see is that these studies are not designed well. They are very short-term, when you think about any possible effects. They are truncating these studies. If you don’t see the effects, then they are concluding that these are safe, which is, in my opinion and many other people’s opinions, irresponsible.”

Oya Yazgan
Studies concluding GMOs are safe often have been conducted by industry-sponsored researchers. Independent researchers have an opposite view.
“A lot of publications and news reports and everything that I look at basically has ties to industry,” Yazgan said. “This is a huge industry — everyone is aware of that — and the feeling is that this is being pushed before we have definitive answers about their safety. That is my concern and my frustration about this as well.”
GMO foods are clearly labeled as such in the European Union. In the U.S., food is either organic or it’s not.
“There is that push because industry has a stronger hold on scientific research and the publications and what’s being made available to the public,” Yazgan said. “In Europe there are more regulations controlling the release of these GMOs and any other substance as well. There is more public support in Europe. There is more business support in the U.S. That’s the biggest difference.”
What’s the best option for concerned consumers? Right now that would be organic, because GMOs aren’t labeled. Big agriculture is trying to wiggle its way out of regulations, Yazgan said.
“The latest technique that is used to make modifications in the genes, they are little different from the previous ones and they do not leave a mark on the DNA of the organisms they are changing,” she said. “The FDA does not consider that genetically engineered, even though they are. They are trying to avoid the regulations.”
Intestinal problems, like irritable bowel syndrome, are on the rise, but not definitively linked to GMOs.
“We need to take a very careful look at these before we play with people’s health,” Yazgan said.
[“source=asunow”]